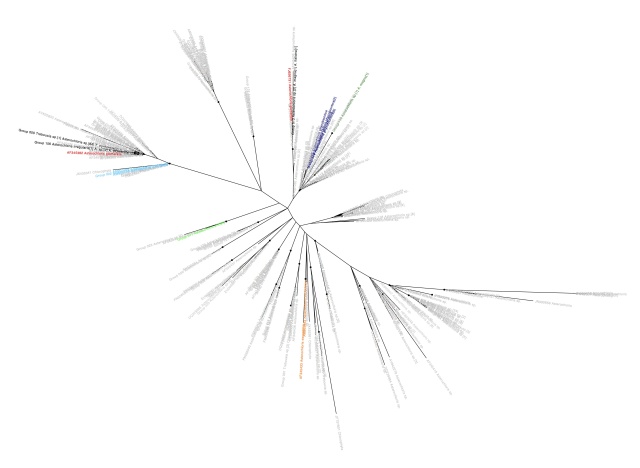
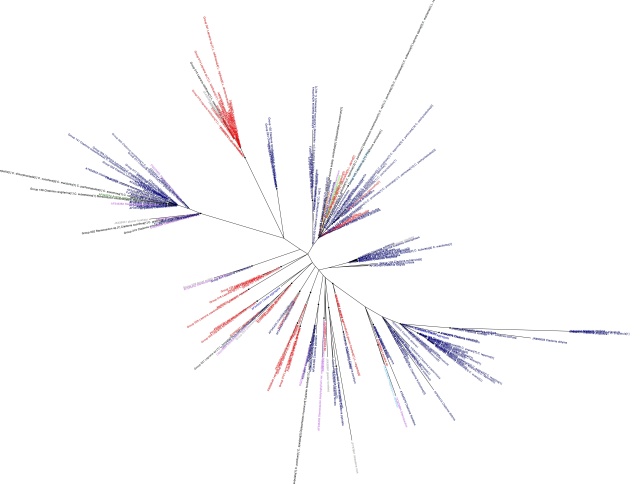
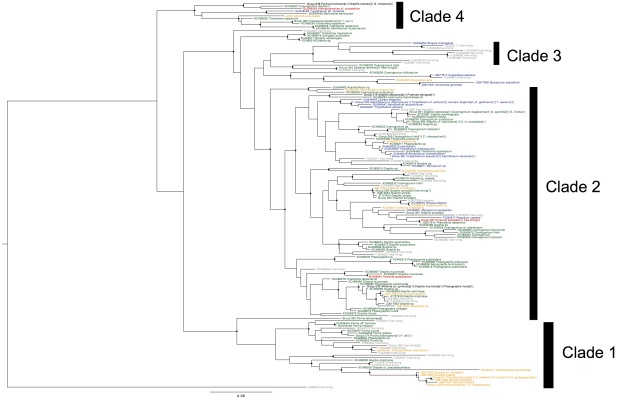
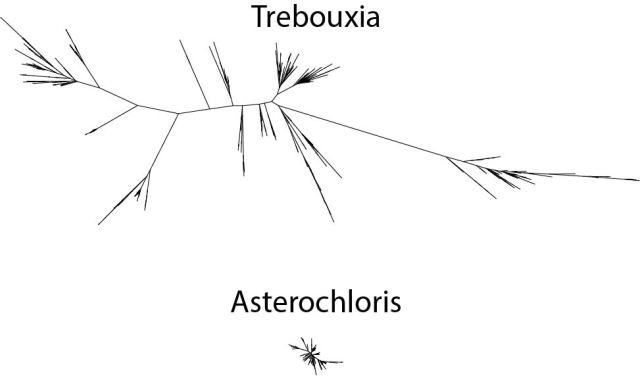
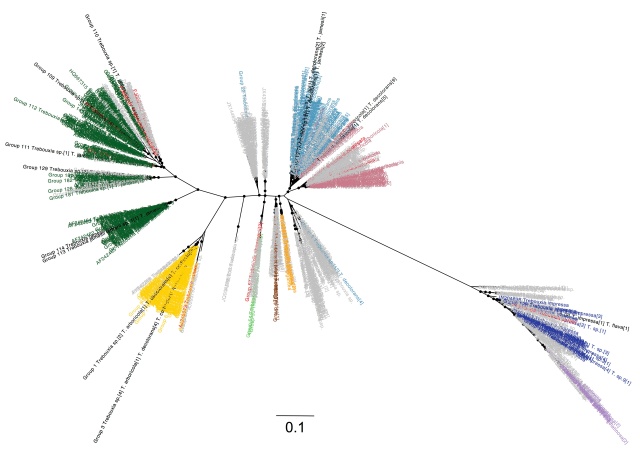
Having gone through the steps to build a phylogeny of the most common lichen photobiont, Trebouxia in my last post, I will now go on to discussing the host association patterns that it reveals. Here is the Trebouxia ITS tree generated previously:
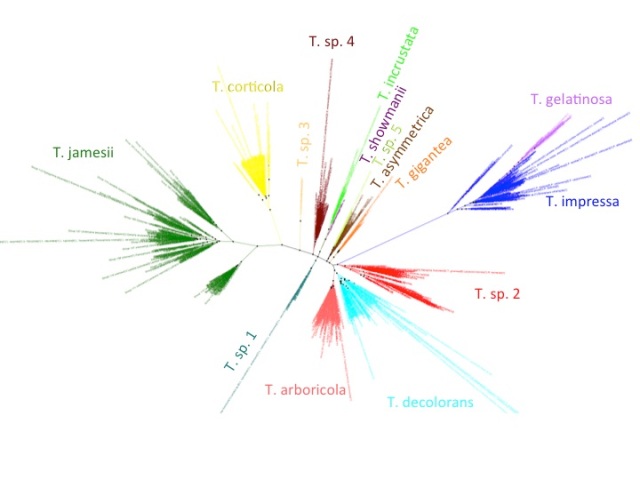
Trebouxia ITS phylogeny. Major clades are differentilly coloured and named according to authentic strains
I’ve coloured all of the taxa within clades according to the colours of the named strains and I’ve assigned unique colours to each clade that does not contain named strains. I have not attempted to break up T. jamesii or T. impressa into sub-clades, though doing so would probably be justified. This will be a topic for a future post. I should also point out that T. jamesii is referred to as T. simplex is some papers.
In contrast to Nostoc photobionts where the fasta headers were consistently labeled with the host information, these sequences are …not. I used a bioperl wrapper to NCBI’s Eutils interface to download genbank format sequences and parsed them to extract host association information from the “host”, “note” and “isolation source” annotions. I also extracted information about the author of each sequence and where it was published while I was at it:
../Scripts/GetGB.pl Trebouxia_ITS_acc.txt heath.obrien-at-gmail-dot-com > Trebouxia_ITS.gbk
../Scripts/ParseHost.pl Trebouxia_ITS.gbk > host_info.txt
../Scripts/GetRef.py Trebouxia_ITS.gbk >ref_info.txt
This information was added to Trebouxia_ITS_metadata.txt in Excel and missing values were filled in manually where possible. I also added culture collection numbers where available and started to fill in locality information, but I haven’t gotten very far with the latter.
Next, I added information about which clade each sequence fell into to the Trebouxia metadata file. The ETE python toolkit was invaluable for this step and was my main proximate motivation for switching from perl to python for my scripting, but I was also really, REALLY tired of having to keep my ‘\@’s and ‘%{$’s straight:
../Script/GetClades.py -t Trebouxia_ITS.nwk -m Trebouxia_ITS_metadata.txt >temp
mv temp Trebouxia_ITS_metadata.txt
In this case, the information was added to the metadata file automatically.
Lastly, I wrote a script to count the number of times that Trebouxia from each clade was associated with each lichen genus that has been sampled:
../Scripts/CountAssociations.py -m Trebouxia_ITS_metadata.txt > AssocaitionCounts.txt
After some fiddling with conditional formatting in Excel, these analyses produced this:

Counts of associations between Trebouxia clades and lichen genera. Colour coding matches the phylogeny
Before discussing the patterns, I should point out that these counts are of how many sequence records have been deposited in genbank, not the number os specimens that have been sampled as many authors deposit representative haplotype sequences rather than all of their data. Incorporating this information will change the counts dramatically in some cases.
The most extensively sampled genus is Letharia, which is exclusively associated with Trebouxia jamesii. In fact, there appears to be strong reciprocal specificity acting between species of Letharia and subclades within T. jamesii, a topic I would like to explore further in the future.
Eight other genera in the Parmeliaceae are also exclusively associated with T. jamesii photobionts, including Cetraria (76 sequences), Evernia (19 sequences), Flavocetraria (17 sequences), Hypogymnia (12 sequences) and Pseudevernia (10 sequences). However, Parmelia (10 sequences), Flavoparmelia (6 sequences), and 4 other genera associate with photobionts in the T. impressa/T. gelatinosa clade (among others) and Parmotrema is highly specific for T. corticola photobionts, with 135 P. tinctorum photobionts grouping with T. corticola (the Trebouxia sp. 3 photobiont is from a different Parmotrema species). The P. tinctorum / T. corticola association is another case of reciprocal specificity as 135 of 141 T. corticola sequences are from P. tinctorum photoboints.
All photobionts from Lasallia (28 sequences) and most from Umbilicaria (105 of 131 sequences) also grouped with T. jamesii, with most of the exceptions being specimens collected in the Antarctic (see also this paper). T. jamesii was also the predominant photobiont of Thamnolia (22 out of 28 sequences) and Chaenotheca (7 of 8 sequences). T. jamesii was also a common photobiont of Lecanora and Lecidea, but photobiont diversity in both of these genera, and the Lecaonraceae in general, is extremely high, with Lecanora photobionts falling into 11 different species and Lecidea photobionts falling into 8. Indeed, 6 species of photobiont have been recovered from L. rupicola alone. Lichens in the Lecanoraceae are the hosts for the vast majority of T. asymmetrica, T. incrustata, T. showmanii and T. sp. 1 photobiots that have been sampled.
Similar to the Parmeliaceae, most of the genera in the Physciaceae are specific for the same Trebouxia lineage, while other genera do not associate with it at all. Physconia (40 sequences) and Phaeophyscia (7 sequences) are exclusively associated with T. impressa, as are 29 out of 36 Physcia sequences while only 2 of 5 Rinodina photobionts and none from Anaptychia (8 sequences) group with T. impressa.
All but 9 of 133 Xanthoria photobionts belong to T. decolorans or T. arboricola, which are sister species. These photobionts also predominate in most of the other genera in the Teloschistaceae, including 53 of 65 Caloplaca photobionts, 4 of 4 Huea photobionts, and 34 of 68 Tephromela photobionts, a genus that is also frequently associated with Trebouxia sp. 2 photobionts.
Specificity is also high for Ramalina, with 139 of 150 photobiont sequences restricted to two Trebouxia clades. Boreoplaca is associated with one of the same photobionts (Trebouxia sp. 2), but not with the most common Ramalina photobiont (T. decolorans; 116 of 150 sequences).
Thus, there is a wide range of association patterns, from extreme reciprocal specificity (135 of 141 T. corticola sequences associated with Parmotrema and all P. tinctorum specimens associated with T. corticola) to generalism (Lecanora rupicola associating with 6 different Trebouxia species). There is some evidence of phylogenetic inertia, as lichen genera from the same family are more likely to share similar photobiont association patterns than unrelated lichens, but there is also a lot of plasticity. There are a lot of ideas out there about the ecological and life history factors that could be causing these differences, but given the complexity of the patterns and our lack of knowledge of things like lichen demography and dispersal mechanisms, it will probably be some time before definitive explanations can be provided.
Heath O’Brien (2013). A preliminary look at host association patterns in Trebouxia PhotobiontDiversity.wordpress.com : http://dx.doi.org/10.6084/m9.figshare.711786